Colistin resistance in Acinetobacter baumannii
Authors: Katarina Novović1 and Branko Jovčić1,2
1 Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, Belgrade, Serbia
2 Faculty of Biology, University of Belgrade, Belgrade, Serbia
E-mails: katarinanovovic@imgge.bg.ac.rs; bjovcic@bio.bg.ac.rs
Reviewer: Younes Smani
Molecular Biology and Biochemical Engineering Dept., Pablo deOlavide University, Seville, Spain (ysma@upo.se)
Colistin resistance in Acinetobacter baumannii
Multidrug-resistant (MDR) and extensively drug-resistant (XDR) A. baumannii are of major public health concern as they cause a high mortality rate in patients. MDR and XDR phenotypes arise from a plethora of intrinsic resistance mechanisms, dynamic gene flow and horizontal gene transfer that shape the variable, open resistome of A. baumannii (Kyriakidis et al., 2020). Colistin is considered a last-line antibiotic with significant activity against MDR and XDR strains and is used as a rescue therapy (Pormohammad et al., 2020). Due to the increasing and non-optimal use of colistin in treating infections, a rapid spread of colistin resistance in A. baumannii has been observed (Pormohammad et al., 2020, Novović and Jovčić, 2023). Published data show that the prevalence of colistin resistance is higher in Southeast Asia and Eastern Mediterranean countries than in other regions of the world, with an overall figure of 11.2% (Pormohammad et al., 2020, Novović and Jovčić, 2023).
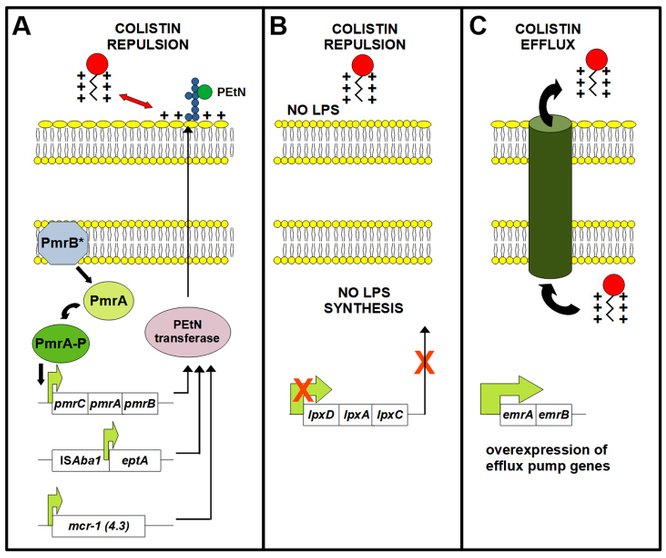
Figure 1. Schematic representation of general mechanisms of colistin resistance in A. baumannii. (A). Addition of phosphoethanolamine (PEtN) moiety to lipid A component of LPS changes the charge of the membrane and results in repulsion of colistin and resistance. This is mostly promoted by altered PmrB histidine kinase that activates expression of the pmrCAB resulting in overexpression of the PmrC PEtN transferase. This could also be achieved by overexpression of EptA or Mcr PEtN transferases. (B). LPS deficiency mediated via the lpxACD genes inactivation results in change of the cell surface charge and repulsion of colistin, as well. (C). Efflux of colistin mediated by efflux pumps has been described as mechanism contributing the resistance to colistin in A. baumannii.
Mechanisms of colistin resistance
The resistance of A. baumannii to colistin is essentially based on two mechanisms, which depend on the complete loss or alteration of the primary target of colistin, the lipid A component of LPS. Both mechanisms lead to a loss of colistin activity due to a decrease of negative charge on the cell surface (Olaitan et al., 2014) (Figure 1A and B). LPS deficiency results from the inactivation of genes encoding enzymes that catalyze the first steps of lipid A biosynthesis (lpxA, lpxC and lpxD genes) (Moffatt et al., 2010). Modification of LPS by the addition of phosphoethanolamine (PEtN) moieties to lipid A is by far the most common mechanism of colistin resistance in A. baumannii. Genetic determinants of this modification are either the chromosomal pmrCAB operon and eptA gene-encoded enzymes or plasmid-encoded mcr genes (Figure 1A). The amino acid changes in the proteins of the PmrAB two-component system that cause colistin resistance are very frequent and diverse, particularly in the histidine kinase A (HisKA) domain of the PmrB protein (Novović and Jovčić, 2023). These changes within the PmrAB proteins lead to overexpression of the phosphoethanolamine transferase PmrC, a key catalyst of PEtN modification of LPS (Arroyo et al., 2011). Increased expression of the eptA gene (pmrC homolog) also enables resistance of A. baumannii to colistin, but is less common (Lesho et al., 2013). Plasmid-encoded mcr genes (especially mcr-1 and mcr-4.3 variants) are the main cause of the rapid spread of resistance to colistin in A. baumannii. Colistin efflux (Figure 1C) (Lin et al., 2017), loss of non-Lpx proteins (lipoproteins) involved in outer membrane (OM) assembly and maintenance (Lima et al., 2017, Bailing et al., 2020) and Omp25 expression reduction have also been described as mechanisms of resistance to colistin (Parra Millán et al., 2018). Heteroresistance to colistin in A. baumannii is based on the same mechanisms as colistin resistance (lpxACD, pmrCAB and efflux pumps) (Novović and Jovčić, 2023).
Global epidemiology of colistin-resistant A. baumannii
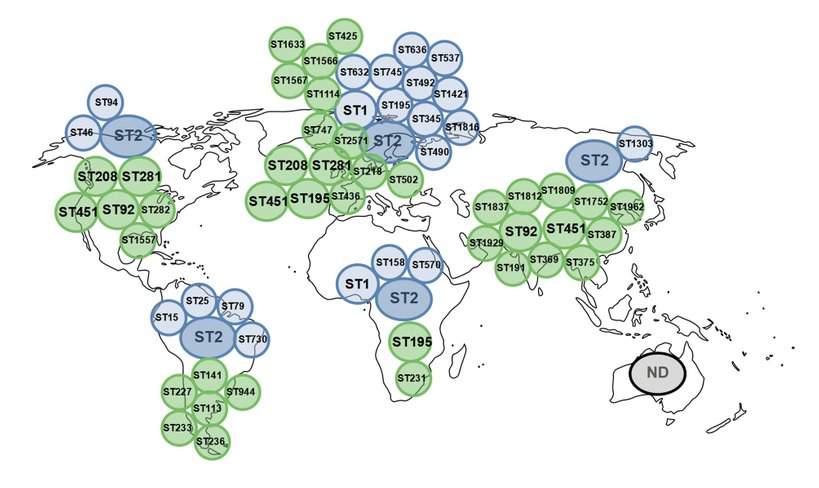
The global distribution of colistin-resistant A. baumannii strains is a picture that will change in the future as we collect more and more data. However, at this stage, using the Pasteur/Oxford MLST scheme to differentiate sequence types (STs), we have identified a predominance of ST2 (Pasteur) worldwide (except Australia) and ST1 (Pasteur) in Europe and Africa, which are not only nosocomial, but also of animal origin (Novović and Jovčić, 2023) (Figure 2). A number of STs (Pasteur) could be considered exclusive to the specific regions: (i) ST195, ST345, ST490, ST492, ST537, ST632, ST636, ST745, ST1421 and ST1816 in Europe, (ii) ST1303 in Asia, (iii) ST158 and ST570 in Africa, (iv) ST46 and ST94 in North America, (v) ST15, ST25, ST79 and ST730 in South America (Novović and Jovčić, 2023) (Figure 2). The more discriminant Oxford MLST scheme showed several STs distributed over different continents, such as ST92 in Asia and North America, ST195 in Europe and Africa, ST208 and ST281 in Europe and North America, ST451 in Europe, Asia and North America (Novović and Jovčić, 2023) (Figure 2). These data indicate a remarkable distribution of colistin-resistant A. baumannii in Europe, Asia and North America.
Figure 2. Global epidemiology of colistin-resistant A. baumannii STs based on Pasteur (blue circles) and Oxford (green circles) MLST scheme. ND – no data available. STs that were found in more than one continent are within circles bigger in size.
References
Kyriakidis I, Vasileiou E, Pana ZD, Tragiannidis A. Acinetobacter baumannii antibiotic resistance mechanisms. Pathogens. 2021 Mar 19;10(3):373.
doi: 10.3390/pathogens10030373
PMID: 33808905
Pormohammad A, Mehdinejadiani K, Gholizadeh P, Nasiri MJ, Mohtavinejad N, Dadashi M, Karimaei S, Safari H, Azimi T. Global prevalence of colistin resistance in clinical isolates of Acinetobacter baumannii: A systematic review and meta-analysis. Microb Pathog. 2020 Feb;139:103887.
doi: 10.1016/j.micpath.2019.103887
PMID: 31765766
Novović K, Jovčić B. Colistin resistance in Acinetobacter baumannii: Molecular mechanisms and epidemiology. Antibiotics (Basel). 2023 Mar 4;12(3):516.
doi: 10.3390/antibiotics12030516
PMID: 36978383
Olaitan AO, Morand S, Rolain JM. Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria. Front Microbiol. 2014 Nov 26;5:643.
PMID: 25505462
Moffatt JH, Harper M, Harrison P, Hale JD, Vinogradov E, Seemann T, Henry R, Crane B, St Michael F, Cox AD, Adler B, Nation RL, Li J, Boyce JD. Colistin resistance in Acinetobacter baumannii is mediated by complete loss of lipopolysaccharide production. Antimicrob Agents Chemother. 2010 Dec;54(12):4971-7.
doi: 10.1128/AAC.00834-10
PMID: 20855724
Arroyo LA, Herrera CM, Fernandez L, Hankins JV, Trent MS, Hancock RE. The pmrCAB operon mediates polymyxin resistance in Acinetobacter baumannii ATCC 17978 and clinical isolates through phosphoethanolamine modification of lipid A. Antimicrob Agents Chemother. 2011 Aug;55(8):3743-51.
doi: 10.1128/AAC.00256-11
PMID: 21646482
Lesho E, Yoon EJ, McGann P, Snesrud E, Kwak Y, Milillo M, Onmus-Leone F, Preston L, St Clair K, Nikolich M, Viscount H, Wortmann G, Zapor M, Grillot-Courvalin C, Courvalin P, Clifford R, Waterman PE. Emergence of colistin-resistance in extremely drug-resistant Acinetobacter baumannii containing a novel pmrCAB operon during colistin therapy of wound infections. J Infect Dis. 2013 Oct 1;208(7):1142-51.
PMID: 23812239
Lin MF, Lin YY, Lan CY. Contribution of EmrAB efflux pumps to colistin resistance in Acinetobacter baumannii. J Microbiol. 2017 Feb;55:130-6.
doi: 10.1007/s12275-017-6408-5
PMID: 28120193
Lima WG, Alves MC, Cruz WS, Paiva MC. Chromosomally encoded and plasmid-mediated polymyxins resistance in Acinetobacter baumannii: A huge public health threat. Eur J Clin Microbiol Infect Dis. 2018 Jun;37:1009-19.
doi: 10.1007/s10096-018-3223-9
PMID: 29524060
Bailing Z, Jieling Z, Honglang L. Mechanism of colistin resistance to Acinetobacter Baumannii and its progress: A review article. Biomed J Sci Tech Res. 2020 Jul;29:22183-8.
doi: 10.26717/BJSTR.2020.29.004755
Parra-Millán R, Vila-Farrés X, Ayerbe-Algaba R, Varese M, Sánchez-Encinales V, Bayó N, Pachón-Ibáñez ME, Teixidó M, Vila J, Pachón J, Giralt E. Synergistic activity of an OmpA inhibitor and colistin against colistin-resistant Acinetobacter baumannii: mechanistic analysis and in vivo efficacy. J Antimicrob Chemother. 2018 Dec 1;73(12):3405-12.
doi: 10.1093/jac/dky343
PMID: 30188994