Antibiotic Resistance
Authors: Benjamin Havenga and Wesaal Khan
Department of Microbiology, Faculty of Science, Stellenbosch University, Stellenbosch, South Africa (18339697@sun.ac.za; wesaal@sun.ac.za)
Reviewers: Gabriela Jorge Da Silva1 and Laurent Poirel2
1Faculty of Pharmacy, University of Coimbra, Coimbra, Portugal (gjsilva@ci.uc.pt)
2Department of Medicine, University of Fribourg, Fribourg, Switzerland (laurent.poirel@unifr.ch)
Acinetobacter baumannii resistome
Ampicillin-sulbactam, imipenem-cilastatin, meropenem, doripenem, amikacin, tobramycin, tigecycline, minocycline, doxycycline, polymyxin E (colistin) and polymyxin B, are generally used for the treatment of A. baumannii infections (Vázquez-López et al., 2020). Yet, fewer effective antibiotic options are practically available as the trends of antibiotic resistance continue to rise among clinical isolates of A. baumannii. In fact, this opportunistic pathogen exhibits intrinsic resistance to several antibiotics (ampicillin, amoxicillin, aztreonam, ertapenem, trimethoprim, chloramphenicol and fosfomycin) (Clinical and Laboratory Standards Institute, 2021) and can accumulate acquired resistance to additional antibiotics (such as β-lactams, aminoglycosides, quinolones, tetracycline, and polymyxins), through vertical acquisition of chromosomal mutations or horizontal gene transfer (Lee et al., 2017). Currently, more A. baumannii strains are turning into multidrug-resistant (MDR), defined as non-susceptible to at least one agent in three or more antimicrobial classes (aminoglycosides, carbapenems, fluoroquinolones, penicillins + β-lactamase inhibitors, extended-spectrum cephalosporins, folate pathway inhibitors, penicillins + β-lactamase inhibitors, polymyxins and tetracyclines), extensively drug-resistant (XDR), defined as non-susceptible to at least one agent in all but two or fewer antimicrobial categories, or pandrug-resistant (PDR), defined as non-susceptible to any agent in all antimicrobial categories (Magiorakos et al., 2011). Such MDR, XDR and PDR strains of A. baumannii employ a variety of enzymatic and non-enzymatic mechanisms to facilitate resistance to antibiotics (Table 1 and Figure 1).
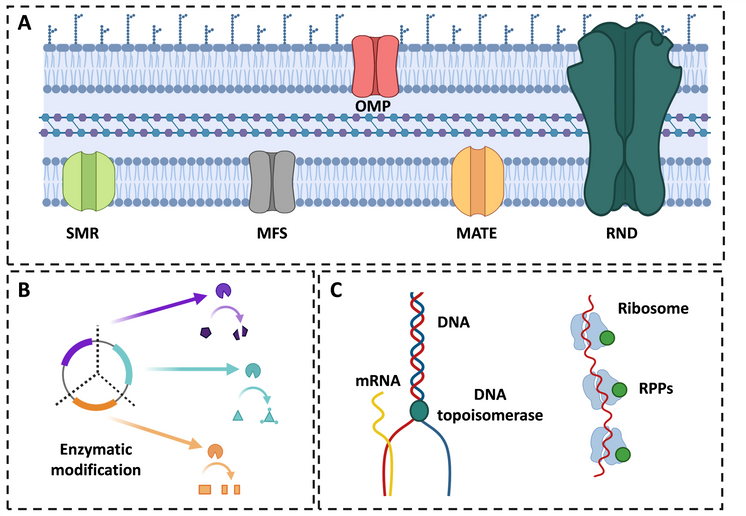
Figure 1. Primary Acinetobacter baumannii antibiotic resistance mechanisms. A) non-enzymatic resistance mechanisms, including efflux pumps such as the outer membrane porin (OMP), small multidrug resistance (SMR), major facilitator super family (MFS), multiple antibiotic and toxin extrusion (MATE) and resistance-nodulation-division (RND); B) enzymatic resistance mechanisms e.g., Ambler class A, B, C and D; C) Alteration of target sites such as DNA topoisomerase, gyrase and ribosomal protection proteins (RPPS) (Adapted from Abdi et al., 2020 and Kyriakidis et al., 2021).
β-lactams
Production of β-lactamases is the main mechanism of resistance to β-lactams and the most prevalent enzymatic antibiotic resistance mechanism employed by A. baumannii (Bush, 2018), where members of all the four Ambler classes (A, B, C and D) (Table 1) have been detected (Lowe et al., 2018). Inherent to all A. baumannii is the Ambler class C (AmpC) cephalosporinase encoded by the ampC gene (Bou & Martínez-Beltrán, 2000). Increased expression of ampC, conferring resistance to broad-spectrum third generation cephalosporins (e.g., ceftazidime and cefotaxime), is mainly mediated by the acquisition of an upstream insertion sequence (IS) element known as ISAba1 (Héritier et al., 2006; Karah et al., 2017). Some variants of the AmpC cephalosporinase, known as Acinetobacter-derived cephalosporinase (ADC), can also facilitate resistance to penicillins, extended-spectrum cephalosporins (cefepime), monobactam (aztreonam) and β-lactamase inhibitors (sulbactam) (Rodríguez-Martínez et al., 2010; Ingti et al., 2020) (Table 1). Apart from the inherent AmpC cephalosporinase, all A. baumannii isolates possess an intrinsic gene encoding an Ambler class D oxacillinase-51-like (OXA-51) enzyme (Héritier et al., 2005; Peleg et al., 2008). OXA-51-like is capable of hydrolyzing penicillins (benzylpenicillin, ampicillin, ticarcillin and piperacillin) and carbapenems (imipenem and meropenem), usually at low levels, when the corresponding gene is overexpressed (Figueiredo et al., 2009a; Figueiredo et al., 2009b; Gordon & Wareham, 2010). Additionally, more than 400 non-intrinsic or acquired oxacillinase enzymes have so far been identified in A. baumannii; of which those who possess carbapenemase activities have currently been clustered into five subgroups: OXA-23, OXA-40, OXA-58, OXA-143 and OXA-235 (Higgins et al., 2013) (Table 1). These carbapenem-hydrolyzing class D β-lactamases (CHDLs) hydrolyze penicillins at high level, and carbapenems weakly. Conversely, they do not significantly hydrolyze broad-spectrum cephalosporins.
Aminoglycosides
Production of aminoglycoside modifying enzymes (AMEs) is the main aminoglycoside resistance mechanism in A. baumannii (Kishk et al., 2021). Based on their mode of action, AMEs have been divided into three groups, the acetyltransferases (AACs), nucleotidyltransferases (ANTs) and phosphotransferases (APHs) (Ramirez & Tolmasky, 2010) (Table 1). Sheikhalizadeh et al. (2017) detected AMEs (ANT, APH and AAC) amongst 94% (n = 89/94) of clinically derived A. baumannii, with ant(2’)-Ia conferring non-susceptibility to tobramycin, kanamycin, amikacin, gentamicin; aph(3’)-VIa conferring non-susceptibility to amikacin and tobramycin; aac(3’)-Ia conferring non-susceptibility to amikacin and tobramycin; and aac(3’)IIa conferring non-susceptibility to kanamycin (Table 1). More recently, Chen et al. (2022) detected the aph(3’)-I, ant(3″)-I and aac(6')-Ib genes amongst 86.32%, 30.53%, and 26.32% A. baumannii strains (n = 92) isolated from an intensive care unit in China. In addition to the above-mentioned enzymatic mechanisms, ribosomal modification (mediated by 16S rRNA methylases or RMTases) and overexpression of efflux pumps [resistance-nodulation-division (RND) or multiple antibiotic and toxin extrusion (MATE) superfamily] have also been associated with aminoglycoside (amikacin, gentamicin, kanamycin, and tobramycin) resistance in A. baumannii (Jouybari et al., 2021) (Table 1 and Figure 1). Currently, the increasing occurrence of acquired RMTases, leading to pan-aminoglycoside resistance, is a concerning phenomenon in A. baumannii and to date 11 RMTases genes have been detected, including armA, rmtA to rmtH, npmA and npmB, with the armA gene being the most prevalent amongst A. baumannii (Taylor et al., 2022).
Quinolones
Quinolone resistance in A. baumannii is primarily mediated by mutations in the quinolone resistance-determining regions of the DNA gyrase subunit A (gyrA) and topoisomerase IV subunit A (parC) genes (Figure 1) (Vila et al., 1995; Vila et al., 1997). In general, a double mutation in both gyrA and parC is needed to acquire high-levels of quinolone resistance (Vila et al., 1997; Hamouda & Amyes, 2004). A. baumannii isolates with triple mutations, involving the gyrA, gyrB and parC genes, exhibit higher ciprofloxacin resistance in comparison to isolates with double mutations in gyrA and parC (Park et al., 2011). Apart from this stepwise target modification mechanism mediated by gyrA, gyrB and parC mutations, efflux systems [RND (including AdeABC, AdeFGH and AdeIJK), MATE or Small Multidrug Resistance (SMR)] have also been associated with quinolone resistance in this pathogen (Yoon et al., 2013; Lee et al., 2017) (Table 1 and Figure 1). The AbeM pump (H+-coupled pump), belonging to the MATE family transporters, has been detected amongst A. baumannii isolates, contributing to quinolone, gentamycin, kanamycin, erythromycin, chloramphenicol, and trimethoprim resistance (Su et al., 2005). The novel SMR efflux system AbeS has also been shown to contribute to quinolone (ciprofloxacin and norfloxacin) resistance in A. baumannii, including the extensively characterised A. baumannii AYE reference strain (Srinivasan et al., 2009).
Tetracycline and glycylcyclines
Tetracycline resistance is primarily facilitated through three main mechanisms, namely efflux pumps, enzyme inactivation and ribosomal protection proteins (RPPs) (Figure 1) (Gordon & Wareham, 2010). The major facilitator super family (MFS) and RND efflux pump systems have been associated with tetracycline resistance (Figure 1). TetA and TetB (MFS family) have frequently been detected in A. baumannii, conferring resistance to tetracycline, doxycycline, and minocycline (Wang et al., 2017) (Table 1). The RND family, including AdeABC and AdeIJK, have also been observed to contribute to tetracycline and glycylcycline (tigecycline) resistance in A. baumannii (Foong et al., 2020). In addition, a synergistic interaction between TetA and the RND-type transporters AdeABC and/or AdeIJK, resulting in higher tetracycline resistance, was reported in the A. baumannii reference strain AYE (Foong et al., 2020). Tetracycline resistance via ribosomal protection, mediated by Tet(M), was also detected in one A. baumannii clinical isolate (Ribera et al., 2003). He et al. (2019) reported on the detection and identification of two plasmid-mediated tigecycline resistance genes, tet(X3) and tet(X4), in various Enterobacteriaceae spp. as well as in A. baumannii (Table 1). Subsequently, Wang et al., (2019) investigated the occurrence of the plasmid-mediated tigecycline resistance genes in clinical A. baumannii isolates (n = 71), resulting in the detection of the novel plasmid-mediated tet(X5) gene. In addition, A. baumannii has been found to harbor the tet(X6) gene, which was detected in strains collected from humans and animal samples in both China and Taiwan (Chen et al., 2021; Hsieh et al., 2021).
Polymyxins
The global emergence of MDR A. baumannii has led to a resurgence in the use of last-resort antibiotics, including polymyxin B and colistin (polymyxin E) (Chen et al., 2019). Polymyxins are usually used as a second line only to treat infections caused by MDR Gram-negative bacteria, such as carbapenem-resistant Enterobacterales, Pseudomonas aeruginosa and A. baumannii (Ledger et al., 2022; Paul et al., 2022). In contrast, they have been extensively used in veterinary medicine for treatment and prevention of infectious diseases and as growth promoters (Poirel et al., 2017). Polymyxins destabilize and disrupt both the outer and cytoplasmic membranes of bacterial cells via direct interaction with the negatively charged phosphate groups of the lipid A component of lipopolysaccharide (LPS) (Chamoun et al., 2021; Ledger et al., 2022). Polymyxin resistance in A. baumannii is mediated by a number of chromosomal and plasmid encoded mechanisms, which can be grouped into: (1) LPS modification; (2) LPS loss (quantitative modification); (3) reduced expression of cofactors involved in LPS synthesis; and (4) downregulation of proteins involved in the export and/or stabilisation of outer membrane precursors (Lima et al., 2018) (Table 1).
LPS modification and LPS loss have been described as the two major mechanisms of polymyxin B and colistin resistance amongst A. baumannii isolates. LPS modification involves mutations in the pmrA and pmrB genes (coding for the PmrAB two-component system), which results in the upregulation and overexpression of the pmrC gene (coding for a pEtN transferase), leading to the addition of phosphoethanolamine (pEtN) to lipid A (Charretier et al., 2018). Additionally, the insertion of ISAba1 upstream of eptA (coding for an alternative PmrC-homolog pEtN transferase) and naxD (encoding an acetyl-galactosamine deacetylase) can cause modification of lipid A and confer colistin resistance in A. baumannii (Lesho et al., 2013; Trebosc et al., 2019; Jovčić et al., 2021). Overexpression of NaxD (acetyl-galactosamine deacetylase), another LPS modifying enzyme regulated by the sensor kinase PmrB, has also been reported to confer colistin resistance (Chin et al., 2015). LPS loss involves mutations in the lipid A biosynthesis genes (Moffatt et al., 2010). Mutations in the lpxA, lpxC, and lpxD genes, or the insertion of ISAba11 in either lpxA or lpxC, have been associated with the complete loss of the LPS (Moffatt et al., 2011; Thi Khanh Nhu et al., 2016; Jovčić et al., 2021).
While colistin resistance was originally related to chromosomal mutations, limiting rapid dissemination (Thi Khanh Nhu et al., 2016), four mobile colistin resistance (mcr) genes, namely mcr-1, mcr-2, mcr-3, and mcr-4.3, have recently been detected in A. baumannii (Khuntayaporn et al., 2022). The mcr genes encode for additional pEtN transferases that can also confer colistin resistance via the addition of pEtN to lipid A (Lima et al., 2018). The mcr-1 gene was detected in one
A. baumannii clinical isolate from Pakistan (Hameed et al., 2019; GenBank accession: MK340994.1). A recent study from Iraq reported on the detection of mcr-1, mcr-2 and mcr-3 among 89, 78 and 82 isolates, respectively, including 66 isolates carrying all the three genes (Al-Kadmy et al., 2020). The mrc-1 gene was also detected in 22 isolates collected from Iraqi hospital settings during 2014 – 2018 (Kareem, 2020). None of the isolates carried mcr-2, mcr-3 or mcr-4 in the latter study. Similarly, mcr-1 was detected in two clinical isolates from Egypt (Shabban et al., 2020), while mcr-1 was reported in one isolate from China (Fan et al., 2020) and three clinical isolates from Pakistan (Ejaz et al., 2021). The mcr-4.3 gene was detected on a plasmid of 25,602 bp (pAB18PR065; GenBank accession: MK360916.1) in an A. baumannii strain isolated from pig fecal sample in China (Ma et al., 2019). Martins-Sorenson et al. (2020) detected the mcr-4.3 gene in a clinical A. baumannii isolate (597A) where it was mediated on plasmid pAb-MCR4.3 (35,502 bp; GenBank accession: CP033872.1). Plasmids pEH_mcr4.3 (18,786-bp; GenBank: CP038261.1) and pEC_mcr4.3 (43,093-bp; GenBank: CP038265.1) were detected in two A. baumannii isolates of human and food origin from the Czech Republic (Bitar et al., 2019). Later, one A. baumannii isolate obtained from frog legs was also found to carry the mcr-4.3 gene (Kalová et al., 2021).
References
Abdi SN, Ghotaslou R, Ganbarov K, Mobed A, Tanomand A, Yousefi M, Asgharzadeh M, Kafil HS. Acinetobacter baumannii efflux pumps and antibiotic resistance. Infect Drug Resist. 2020 Feb 12;13:423-434.
doi: 10.2147/IDR.S228089
PMID: 32104014
Al-Kadmy IMS, Ibrahim SA, Al-Saryi N, Aziz SN, Besinis A, Hetta HF. Prevalence of genes involved in colistin resistance in Acinetobacter baumannii: First report from Iraq. Microb Drug Resist. 2020 Jun;26(6):616-622.
PMID: 31816255
Bitar I, Medvecky M, Gelbicova T, Jakubu V, Hrabak J, Zemlickova H, Karpiskova R, Dolejska M. Complete nucleotide sequences of mcr-4.3-carrying plasmids in Acinetobacter baumannii sequence type 345 of human and food origin from the Czech Republic, the first case in Europe. Antimicrob Agents Chemother. 2019 Sep 23;63(10):e01166-19.
doi: 10.1128/AAC.01166-19
PMID: 31332072
Bou G, Martínez-Beltrán J. Cloning, nucleotide sequencing, and analysis of the gene encoding an AmpC beta-lactamase in Acinetobacter baumannii. Antimicrob Agents Chemother. 2000 Feb;44(2):428-32.
doi: 10.1128/AAC.44.2.428-432.2000.
PMID: 10639377
Bush K. Past and present perspectives on β-lactamases. Antimicrob Agents Chemother. 2018 Sep 24;62(10):e01076-18. doi: 10.1128/AAC.01076-18
PMID: 30061284
Chamoun S, Welander J, Martis-Thiele MM, Ntzouni M, Claesson C, Vikström E, Turkina MV. Colistin Dependence in extensively drug-resistant Acinetobacter baumannii strain is associated with ISAjo2 and ISAba13 insertions and multiple cellular responses. Int J Mol Sci. 2021 Jan 8;22(2):576.
doi: 10.3390/ijms22020576.
PMID: 33430070
Charretier Y, Diene SM, Baud D, Chatellier S, Santiago-Allexant E, van Belkum A, Guigon G, Schrenzel J. Colistin heteroresistance and involvement of the PmrAB regulatory system in Acinetobacter baumannii. Antimicrob Agents Chemother. 2018 Aug 27;62(9):e00788-18.
doi: 10.1128/AAC.00788-18
PMID: 29914966
Chen C, Cui CY, Wu XT, Fang LX, He Q, He B, Long TF, Liao XP, Chen L, Liu YH, Sun J. Spread of tet(X5) and tet(X6) genes in multidrug-resistant Acinetobacter baumannii strains of animal origin. Vet Microbiol. 2021 Feb;253:108954.
doi: 10.1016/j.vetmic.2020.108954
PMID: 33373881
Chen F, Deng X, Wang Z, Wang L, Wang K, Gao L. Treatment of severe ventriculitis caused by extensively drug-resistant Acinetobacter baumannii by intraventricular lavage and administration of colistin. Infect Drug Resist. 2019 Jan 21;12:241-247.
doi: 10.2147/IDR.S186646
PMID: 30718963
Chen ZR, Guo HW, Liu J, Pan Q, Fu MZ, Qiu YK, Wong NK, Huang YC. Resistance traits and molecular characterization of multidrug-resistant Acinetobacter baumannii isolates from an intensive care unit of a tertiary hospital in Guangdong, southern China. Int Microbiol. 2022 Aug;25(3):471-479.
doi: 10.1007/s10123-022-00233-0
PMID: 35098390
Chin CY, Gregg KA, Napier BA, Ernst RK, Weiss DS. A PmrB-Regulated deacetylase required for Lipid A modification and polymyxin resistance in Acinetobacter baumannii. Antimicrob Agents Chemother. 2015 Dec;59(12):7911-4.
doi: 10.1128/AAC.00515-15
PMID: 26459891
Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial disk susceptibility tests. 2021. http://em100.edaptivedocs.net/dashboard.aspx [Accessed: 1 January 2021].
Ejaz H, Younas S, Qamar MU, Junaid K, Abdalla AE, Abosalif KOA, Alameen AAM, Elamir MYM, Ahmad N, Hamam SSM, Salem EHM, Bukhari SNA. Molecular epidemiology of extensively drug-resistant mcr encoded colistin-resistant bacterial strains co-expressing multifarious β-lactamases. Antibiotics (Basel). 2021 Apr 20;10(4):467.
doi: 10.3390/antibiotics10040467
PMID: 33923991
Fan R, Li C, Duan R, Qin S, Liang J, Xiao M, Lv D, Jing H, Wang X. Retrospective screening and analysis of mcr-1 and bla NDM in Gram-negative bacteria in China, 2010–2019. Front Microbiol. 2020 Feb 11;11:121.
PMID: 32117144
Figueiredo S, Poirel L, Croize J, Recule C, Nordmann P. In vivo selection of reduced susceptibility to carbapenems in Acinetobacter baumannii related to ISAba1-mediated overexpression of the natural bla(OXA-66) oxacillinase gene. Antimicrob Agents Chemother. 2009a Jun;53(6):2657-9.
doi: 10.1128/AAC.01663-08
PMID: 19307373
Figueiredo S, Poirel L, Papa A, Koulourida V, Nordmann P. Overexpression of the naturally occurring blaOXA-51 gene in Acinetobacter baumannii mediated by novel insertion sequence ISAba9. Antimicrob Agents Chemother. 2009b Sep;53(9):4045-7.
doi: 10.1128/AAC.00292-09
PMID: 19564367
Foong WE, Wilhelm J, Tam HK, Pos KM. Tigecycline efflux in Acinetobacter baumannii is mediated by TetA in synergy with RND-type efflux transporters. J Antimicrob Chemother. 2020 May 1;75(5):1135-1139.
doi: 10.1093/jac/dkaa015
PMID: 32049277
Gordon NC, Wareham DW. Multidrug-resistant Acinetobacter baumannii: Mechanisms of virulence and resistance. Int J Antimicrob Agents. 2010 Mar;35(3):219-26.
doi: 10.1016/j.ijantimicag.2009.10.024
PMID: 20047818
Hameed F, Khan MA, Muhammad H, Sarwar T, Bilal H, Rehman TU. Plasmid-mediated mcr-1 gene in Acinetobacter baumannii and Pseudomonas aeruginosa: First report from Pakistan. Rev Soc Bras Med Trop. 2019 Sep 5;52:e20190237. doi: 10.1590/0037-8682-0237-2019
PMID: 31508785
Hamouda A, Amyes SG. Novel gyrA and parC point mutations in two strains of Acinetobacter baumannii resistant to ciprofloxacin. J Antimicrob Chemother. 2004 Sep;54(3):695-6.
doi: 10.1093/jac/dkh368
PMID: 15282231
Havenga B, Reyneke B, Waso-Reyneke M, Ndlovu T, Khan S, Khan W. Biological control of Acinetobacter baumannii: In vitro and in vivo activity, limitations, and combination therapies. Microorganisms. 2022 May 19;10(5):1052.
doi: 10.3390/microorganisms10051052
PMID: 35630494
He T, Wang R, Liu D, Walsh TR, Zhang R, Lv Y, Ke Y, Ji Q, Wei R, Liu Z, Shen Y, Wang G, Sun L, Lei L, Lv Z, Li Y, Pang M, Wang L, Sun Q, Fu Y, Song H, Hao Y, Shen Z, Wang S, Chen G, Wu C, Shen J, Wang Y. Emergence of plasmid-mediated high-level tigecycline resistance genes in animals and humans. Nat Microbiol. 2019 Sep;4(9):1450-1456.
doi: 10.1038/s41564-019-0445-2
PMID: 31133751
Héritier C, Poirel L, Fournier PE, Claverie JM, Raoult D, Nordmann P. Characterization of the naturally occurring oxacillinase of Acinetobacter baumannii. Antimicrob Agents Chemother. 2005 Oct;49(10):4174-9.
doi: 10.1128/AAC.49.10.4174-4179.2005
PMID: 16189095
Héritier C, Poirel L, Nordmann P. Cephalosporinase over-expression resulting from insertion of ISAba1 in Acinetobacter baumannii. Clin Microbiolog Infect. 2006 Feb 1;12(2):123-30.
doi: 10.1111/j.1469-0691.2005.01320.x
PMID: 16441449
Higgins PG, Pérez-Llarena FJ, Zander E, Fernández A, Bou G, Seifert H. OXA-235, a novel class D β-lactamase involved in resistance to carbapenems in Acinetobacter baumannii. Antimicrob Agents Chemother. 2013 May;57(5):2121-6.
doi: 10.1128/AAC.02413-12
PMID: 23439638
Hsieh YC, Wu JW, Chen YY, Quyen TLT, Liao WC, Li SW, Chen YC, Pan YJ. An Outbreak of tet(X6)-Carrying Tigecycline-resistant Acinetobacter baumannii isolates with a new capsular type at a hospital in Taiwan. Antibiotics (Basel). 2021 Oct 12;10(10):1239.
doi: 10.3390/antibiotics10101239
PMID: 34680819
Ingti B, Upadhyay S, Hazarika M, Khyriem AB, Paul D, Bhattacharya P, Joshi SR, Bora D, Dhar D, Bhattacharjee A. Distribution of carbapenem resistant Acinetobacter baumannii with blaADC-30 and induction of ADC-30 in response to beta-lactam antibiotics. Res Microbiol. 2020 Apr-Jun;171(3-4):128-133.
doi: 10.1016/j.resmic.2020.01.002
PMID: 31988011
Jouybari MA, Ahanjan M, Mirzaei B, Goli HR. Role of aminoglycoside-modifying enzymes and 16S rRNA methylase (ArmA) in resistance of Acinetobacter baumannii clinical isolates against aminoglycosides. Rev Soc Bras Med Trop. 2021 Jan 29;54:e05992020.
doi: 10.1590/0037-8682-0599-2020
PMID: 33533819
Jovčić B, Novovic K, Dekic S, Hrenovic J. Colistin resistance in environmental Isolates of Acinetobacter baumannii. Microb Drug Resist. 2021 Mar;27(3):328-336.
PMID: 32762604
Kalová A, Gelbíčová T, Overballe-Petersen S, Litrup E, Karpíšková R. Characterisation of colistin-resistant Enterobacterales and Acinetobacter strains carrying mcr genes from asian aquaculture products. Antibiotics (Basel). 2021 Jul 9;10(7):838.
doi: 10.3390/antibiotics10070838
PMID: 34356760
Karah N, Jolley KA, Hall RM, Uhlin BE. Database for the ampC alleles in Acinetobacter baumannii. PLoS One. 2017 May 1;12(5):e0176695.
doi: 10.1371/journal.pone.0176695
PMID: 28459877
Kareem SM. Emergence of mcr-and fosA3-mediated colistin and fosfomycin resistance among carbapenem-resistant Acinetobacter baumannii in Iraq. Meta Gene. 2020 Sep 1;25:100708.
doi: 10.1016/j.mgene.2020.100708
Khuntayaporn P, Thirapanmethee K, Chomnawang MT. An update of mobile colistin resistance in non-fermentative Gram-negative bacilli. Front Cell Infect Microbiol. 2022 Jun 17;12:882236.
doi: 10.3389/fcimb.2022.882236
PMID: 35782127
Kishk R, Soliman N, Nemr N, Eldesouki R, Mahrous N, Gobouri A, Azab E, Anani M. Prevalence of Aminoglycoside resistance and aminoglycoside modifying enzymes in Acinetobacter baumannii among intensive care unit patients, Ismailia, Egypt. Infect Drug Resist. 2021 Jan 19;14:143-150.
doi: 10.2147/IDR.S290584
PMID: 33519215
Kyriakidis I, Vasileiou E, Pana ZD, Tragiannidis A. Acinetobacter baumannii antibiotic resistance mechanisms. Pathogens. 2021 Mar 19;10(3):373.
doi: 10.3390/pathogens10030373
PMID: 33808905
Ledger EVK, Sabnis A, Edwards AM. Polymyxin and lipopeptide antibiotics: Membrane-targeting drugs of last resort. Microbiology (Reading). 2022 Feb;168(2):001136.
doi: 10.1099/mic.0.001136
PMID: 35118938
Lee CR, Lee JH, Park M, Park KS, Bae IK, Kim YB, Cha CJ, Jeong BC, Lee SH. Biology of Acinetobacter baumannii: Pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front Cell Infect Microbiol. 2017 Mar 13;7:55.
PMID: 28348979
Lesho E, Yoon EJ, McGann P, Snesrud E, Kwak Y, Milillo M, Onmus-Leone F, Preston L, St Clair K, Nikolich M, Viscount H, Wortmann G, Zapor M, Grillot-Courvalin C, Courvalin P, Clifford R, Waterman PE. Emergence of colistin-resistance in extremely drug-resistant Acinetobacter baumannii containing a novel pmrCAB operon during colistin therapy of wound infections. J Infect Dis. 2013 Oct 1;208(7):1142-51.
PMID: 23812239
Lima WG, Alves MC, Cruz WS, Paiva MC. Chromosomally encoded and plasmid-mediated polymyxins resistance in Acinetobacter baumannii: A huge public health threat. Eur J Clin Microbiol Infect Dis. 2018 Jun;37(6):1009-1019.
doi: 10.1007/s10096-018-3223-9
PMID: 29524060
Lowe M, Ehlers MM, Ismail F, Peirano G, Becker PJ, Pitout JDD, Kock MM. Acinetobacter baumannii: Epidemiological and beta-lactamase data from two tertiary academic hospitals in Tshwane, South Africa. Front Microbiol. 2018 Jun 12;9:1280. doi: 10.3389/fmicb.2018.01280.
PMID: 29946315
Ma F, Shen C, Zheng X, Liu Y, Chen H, Zhong L, Liang Y, Liao K, Xia Y, Tian GB, Yang Y. Identification of a novel plasmid carrying mcr-4.3 in an Acinetobacter baumannii strain in China. Antimicrob Agents Chemother. 2019 May 24;63(6):e00133-19.
doi: 10.1128/AAC.00133-19
PMID: 30936095
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012 Mar;18(3):268-81.
doi: 10.1111/j.1469-0691.2011.03570.x
PMID: 21793988
Martins-Sorenson N, Snesrud E, Xavier DE, Cacci LC, Iavarone AT, McGann P, Riley LW, Moreira BM. A novel plasmid-encoded mcr-4.3 gene in a colistin-resistant Acinetobacter baumannii clinical strain. J Antimicrob Chemother. 2020 Jan 1;75(1):60-64.
doi: 10.1093/jac/dkz413
PMID: 31578567
Moffatt JH, Harper M, Harrison P, Hale JD, Vinogradov E, Seemann T, Henry R, Crane B, St Michael F, Cox AD, Adler B, Nation RL, Li J, Boyce JD. Colistin resistance in Acinetobacter baumannii is mediated by complete loss of lipopolysaccharide production. Antimicrob Agents Chemother. 2010 Dec;54(12):4971-7.
doi: 10.1128/AAC.00834-10
PMID: 20855724
Park S, Lee KM, Yoo YS, Yoo JS, Yoo JI, Kim HS, Lee YS, Chung GT. Alterations of gyrA, gyrB, and parC and activity of efflux pump in fluoroquinolone-resistant Acinetobacter baumannii. Osong Public Health Res Perspect. 2011 Dec;2(3):164-70. doi: 10.1016/j.phrp.2011.11.040
PMID: 24159468
Paul M, Carrara E, Retamar P, Tängdén T, Bitterman R, Bonomo RA, de Waele J, Daikos GL, Akova M, Harbarth S, Pulcini C, Garnacho-Montero J, Seme K, Tumbarello M, Lindemann PC, Gandra S, Yu Y, Bassetti M, Mouton JW, Tacconelli E, Rodríguez-Baño J. European Society of Clinical Microbiology and Infectious Diseases (ESCMID) guidelines for the treatment of infections caused by multidrug-resistant Gram-negative bacilli (endorsed by European society of intensive care medicine). Clin Microbiol Infect. 2022 Apr;28(4):521-547.
doi: 10.1016/j.cmi.2021.11.025
PMID: 34923128
Peleg AY, Seifert H, Paterson DL. Acinetobacter baumannii: Emergence of a successful pathogen. Clin Microbiol Rev. 2008 Jul;21(3):538-82.
doi: 10.1128/CMR.00058-07
PMID: 18625687
Poirel L, Jayol A, Nordmann P. Polymyxins: Antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev. 2017 Apr;30(2):557-596.
doi: 10.1128/CMR.00064-16
PMID: 28275006
Ramirez MS, Tolmasky ME. Aminoglycoside modifying enzymes. Drug Resist Updat. 2010 Dec;13(6):151-71.
doi: 10.1016/j.drup.2010.08.003
PMID: 20833577
Ribera A, Ruiz J, Vila J. Presence of the TetM determinant in a clinical isolate of Acinetobacter baumannii. Antimicrob Agents Chemother. 2003 Jul;47(7):2310-2.
doi: 10.1128/AAC.47.7.2310-2312.2003
PMID: 12821485
Rodríguez-Martínez JM, Nordmann P, Ronco E, Poirel L. Extended-spectrum cephalosporinase in Acinetobacter baumannii. Antimicrob Agents Chemother. 2010 Aug;54(8):3484-8.
doi: 10.1128/AAC.00050-10
PMID: 20547808
Shabban M, Fahim NA, Montasser K, El Magd NM. Resistance to colistin mediated by mcr-1 among multidrug resistant Gram-negative pathogens at a tertiary care hospital, Egypt. JPAM. 2020 Jun 1;14(2):1125-32.
Sheikhalizadeh V, Hasani A, Ahangarzadeh Rezaee M, Rahmati-Yamchi M, Hasani A, Ghotaslou R, Goli HR. Comprehensive study to investigate the role of various aminoglycoside resistance mechanisms in clinical isolates of Acinetobacter baumannii. J Infect Chemother. 2017 Feb;23(2):74-79.
doi: 10.1016/j.jiac.2016.09.012
PMID: 27889248
Srinivasan VB, Rajamohan G, Gebreyes WA. Role of AbeS, a novel efflux pump of the SMR family of transporters, in resistance to antimicrobial agents in Acinetobacter baumannii. Antimicrob Agents Chemother. 2009 Dec;53(12):5312-6. doi: 10.1128/AAC.00748-09
PMID: 19770280
Su XZ, Chen J, Mizushima T, Kuroda T, Tsuchiya T. AbeM, an H+-coupled Acinetobacter baumannii multidrug efflux pump belonging to the MATE family of transporters. Antimicrob Agents Chemother. 2005 Oct;49(10):4362-4.
doi: 10.1128/AAC.49.10.4362-4364.2005.
PMID: 16189122
Taylor E, Jauneikaite E, Sriskandan S, Woodford N, Hopkins KL. Novel 16S rRNA methyltransferase RmtE3 in Acinetobacter baumannii ST79. J Med Microbiol. 2022 May;71(5).
doi: 10.1099/jmm.0.001531
PMID: 35588089
Thi Khanh Nhu N, Riordan DW, Do Hoang Nhu T, Thanh DP, Thwaites G, Huong Lan NP, Wren BW, Baker S, Stabler RA. The induction and identification of novel colistin resistance mutations in Acinetobacter baumannii and their implications. Sci Rep. 2016 Jun 22;6:28291.
doi: 10.1038/srep28291
PMID: 27329501
Trebosc V, Gartenmann S, Tötzl M, Lucchini V, Schellhorn B, Pieren M, Lociuro S, Gitzinger M, Tigges M, Bumann D, Kemmer C. Dissecting colistin resistance mechanisms in extensively drug-resistant Acinetobacter baumannii clinical isolates. mBio. 2019 Jul 16;10(4):e01083-19.
PMID: 31311879
Vázquez-López R, Solano-Gálvez SG, Juárez Vignon-Whaley JJ, Abello Vaamonde JA, Padró Alonzo LA, Rivera Reséndiz A, Muleiro Álvarez M, Vega López EN, Franyuti-Kelly G, Álvarez-Hernández DA, Moncaleano Guzmán V, Juárez Bañuelos JE, Marcos Felix J, González Barrios JA, Barrientos Fortes T. Acinetobacter baumannii resistance: A real challenge for clinicians. Antibiotics (Basel). 2020 Apr 23;9(4):205.
doi: 10.3390/antibiotics9040205
PMID: 32340386
Vila J, Ruiz J, Goñi P, Marcos A, Jimenez de Anta T. Mutation in the gyrA gene of quinolone-resistant clinical isolates of Acinetobacter baumannii. Antimicrob Agents Chemother. 1995 May;39(5):1201-3.
PMID: 7625818
Vila J, Ruiz J, Goñi P, Jimenez de Anta T. Quinolone-resistance mutations in the topoisomerase IV parC gene of Acinetobacter baumannii. J Antimicrob Chemother. 1997 Jun;39(6):757-62.
doi: 10.1093/jac/39.6.757
PMID: 9222045
Wang L, Liu D, Lv Y, Cui L, Li Y, Li T, Song H, Hao Y, Shen J, Wang Y, Walsh TR. Novel plasmid-mediated tet(X5) gene conferring resistance to tigecycline, eravacycline, and omadacycline in a clinical Acinetobacter baumannii isolate. Antimicrob Agents Chemother. 2019 Dec 20;64(1):e01326-19.
doi: 10.1128/AAC.01326-19
PMID: 31611352
Wang P, McElheny CL, Mettus RT, Shanks RMQ, Doi Y. Contribution of the TetB efflux pump to minocycline susceptibility among carbapenem-resistant Acinetobacter baumannii strains. Antimicrob Agents Chemother. 2017 Sep 22;61(10):e01176-17.
doi: 10.1128/AAC.01176-17
PMID: 28739789
Yoon EJ, Courvalin P, Grillot-Courvalin C. RND-type efflux pumps in multidrug-resistant clinical isolates of Acinetobacter baumannii: Major role for AdeABC overexpression and AdeRS mutations. Antimicrob Agents Chemother. 2013 Jul;57(7):2989-95.
doi: 10.1128/AAC.02556-12
PMID: 23587960